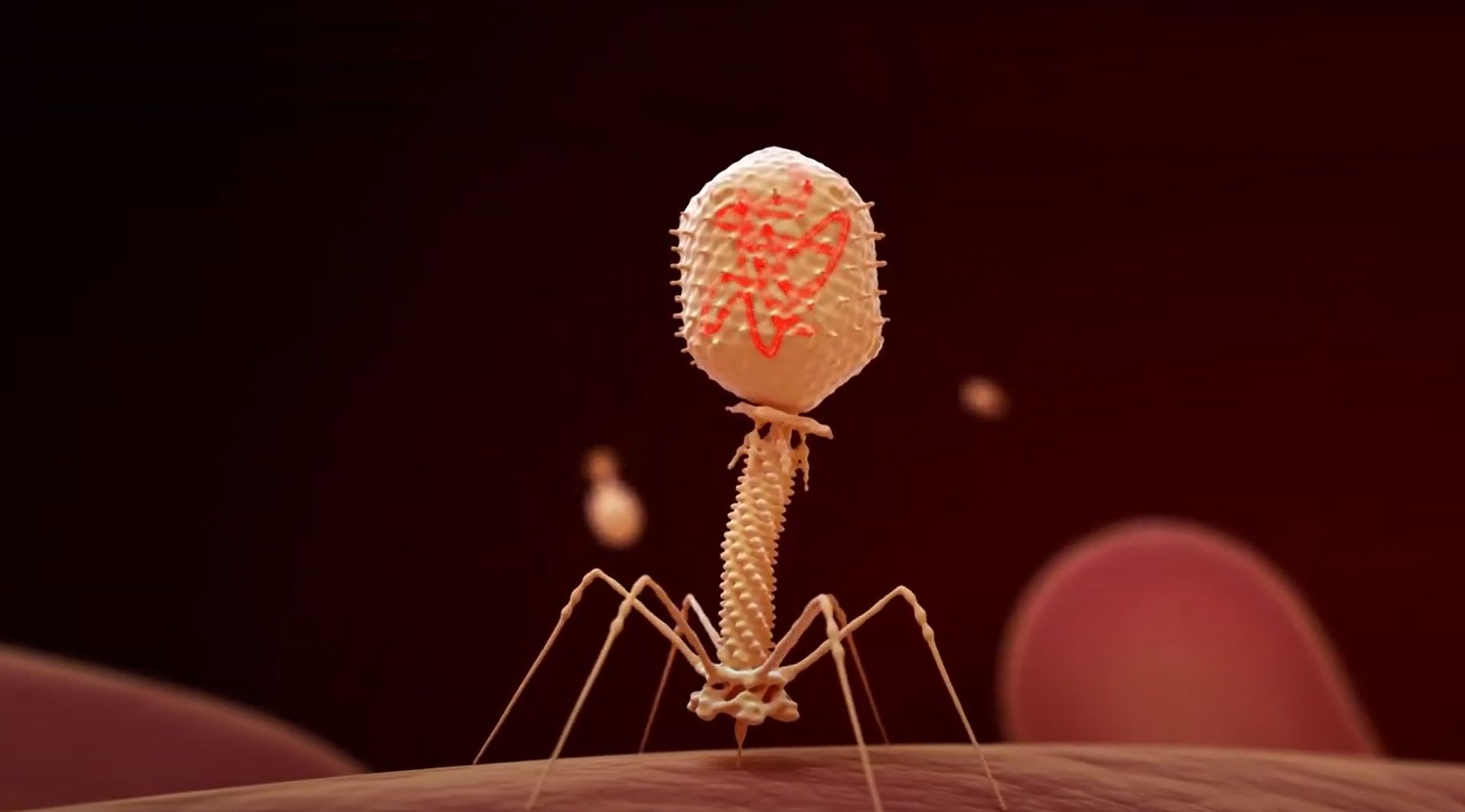
Jill Banfield and her team have used genome-resolved metagenomics to find new CRISPR systems, huge viruses, whole new branches on the tree of life, and more — but what exactly is metagenomics? We wanted to take a step back and introduce readers to the Banfield lab’s pioneering techniques for finding and understanding new microbes, with a little help from Banfield lab Associate Project Scientist Spencer Diamond.
What is metagenomics and what can we learn from it?
Metagenomics is the study of DNA from a mixed community of organisms, usually microbes. It involves taking a sample of microbes from an environment like soil, an aquifer, or part of the body. The genetic material is isolated, and the resulting sample has genetic material from all the community members, mixed together. A lot can be learned from what kind of genes are present in the sample, like what species are there, what metabolic pathways they use, how their immune systems work, and more.
“Only 1% of microbes in the world have been grown in a laboratory,” says Diamond. “And despite our best efforts, most of them still can’t be. Sometimes it’s because we don’t know the exact conditions they need to grow, but it’s also because a lot of microbes are social — they live in symbiotic or parasitic relationships with other microbes, and they can’t grown on their own.”
Metagenomics is the main way to study the many species we can’t grow in labs. These organisms might have unique metabolic pathways, and could be new sources of antibiotics and genome-editing tools like CRISPR.
What is genome-resolved metagenomics?
In genome-resolved metagenomics, the complex mixture of DNA sequences are fragmented, sequenced, and assembled into longer, continuous sequences that represent parts of, or whole, genomes. “Genome-resolved metagenomics,” says Diamond, “gives you the complete picture of every organism who is there, what their genome is, and what they might be able to do.” IGI Microbiology Director Jill Banfield pioneered genome-resolved metagenomics in 2004.
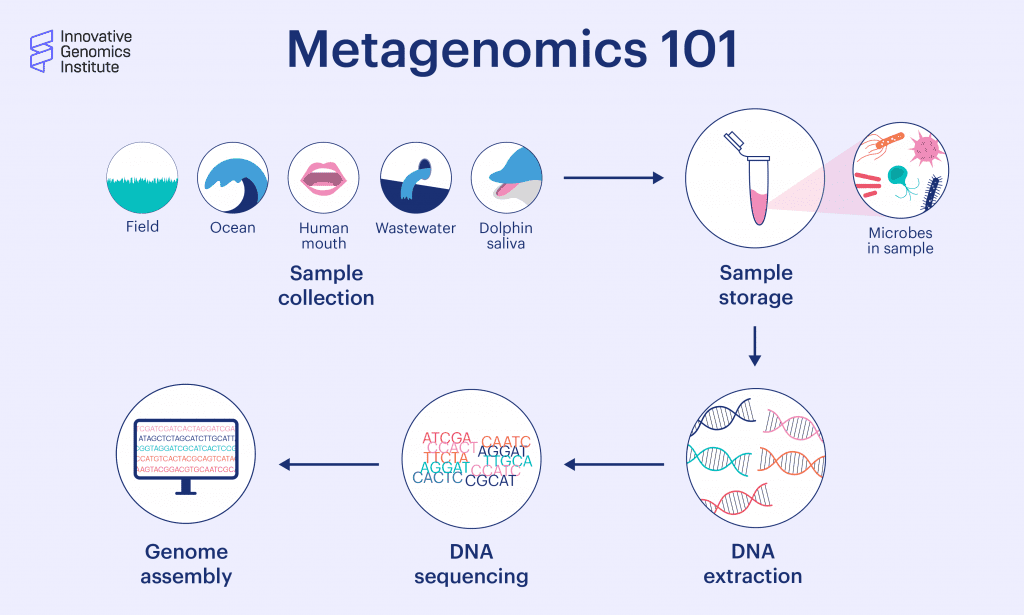
What big findings have come from metagenomics so far?
Genome-resolved metagenomics has been used to find and study organisms living in some of the most remote and extreme environments, like geysers, deserts, and toxic waste water. Some of the most significant findings are new species and whole new classes of organisms that were previously unknown to science.
“Metagenomics has rewritten the tree of life,” says Diamond. “Now we know that there is a massive amount of genetic diversity, and humans and animals are just this little blip in the corner. In an evolutionary sense, we are insignificant and small! It’s kind of ironic because what we’re finding is that most living things are actually extremely small — literally right under our feet is such dramatic genetic diversity. Most of the diversity of life on earth is actually in microorganisms. Metagenomics has found whole huge groups of microorganisms we didn’t know existed before. Many of them are ultrasmall, smaller than we realized was possible.”
What are metatranscriptomics and metaproteomics? What can we learn from them?
“Genomics means, instead of targeting an individual small piece of DNA, you just sequence all of the DNA in a mixture and use a computer to sort out the data,” explains Diamond. “That idea of ‘omics’ was applied to other areas.” Metatranscriptomics is a method of sequencing all of the RNA in a mixed sample, and metaproteomics is a way of identifying proteins in a mixed sample.
“Life has four main macromolecules: proteins, lipids (fats), carbohydrates, and nucleic acids – DNA and RNA,” says Diamond. “And each of those things tells you a piece of the story about what’s going on in an organism. DNA basically tells you who it is and what it could possibly do. What it actually does will vary with circumstances, and mRNA tells you what an organism is actively doing on a small timescale, what genes are being turned on and off. The proteome is a fancy word for all the proteins that are present in that organism — it’s really a ground truth about what proteins are actually present in an organism at a moment in time. Each provides a different view of what’s going on as well as what’s happening on different timescales.”
What are some of the potential applications of metagenomics?

“Metagenomics has been extremely fruitful in the discovery of new molecular biology tools,” says Diamond, “So many genetic techniques are based on matching DNA to things we already know. With metagenomics you don’t know what you’re getting! We can recover DNA and genomes we never knew existed, or even suspected were in a sample. There’s so much more to be discovered in these unknown genomes.”
In fact, CRISPR genome editing technology is an adaptation of CRISPR immune systems housed in the genomes of many bacteria and archaea, and CRISPR-related proteins from different organisms are being developed for editing, diagnostic tools, and more. Microorganisms can also be sources of inspiration for new medicines, like antibiotics, antifungals, and statins. What else might be lurking in mysterious new genomes?
“In terms of the global ecosystem, animal metabolism is pretty boring, we basically just eat food and breathe air. But these tiny organisms can do unbelievable different types of metabolism. They’re responsible for the turnover of all the nutrients in our biosphere,” says Diamond. For example, some microorganisms eat methane, a greenhouse gas, and others convert soil nitrogen into a form plants need for growing. As we learn more about these organisms, we may be able to use their metabolic capabilities to reduce greenhouse gas emissions, improve soil health, reduce farm inputs, capture carbon from the atmosphere, and more.
What’s next frontier for metagenomics?
“We’ve really just been kind of like birdwatchers so far — we go to these environments and we scope out what’s going on. We’ve gotten to the point where we can use metagenomics to identify the organisms and what they might be doing,” says Diamond.
In a new paper, Diamond along with Benjamin Rubin, Brady Cress, and other researchers from the Banfield and Doudna labs take steps towards being able to edit microbial genomes without even having to isolate the organisms out of their native environments and microbial communities. First, they mimicked a soil microbiome by growing a group of nine soil microorganisms together in the lab. Treating the whole microbiome with genome editing reagents, they were able to successfully edit genes in specific organisms only.
Moving closer towards editing in a natural environment, the team started working with a human stool sample, which is rich in microorganisms. From the sample, they grew the organisms together, creating a stable community made mostly of 14 different types of microorganisms. Treating the whole community with genome editing tools, they were able to specifically edit individual E. coli strains, targeting genes that are related to infection.
In the future, community editing approaches might be used to directly edit select organisms in their natural environments to treat human diseases, mitigate climate change, and more. Learn more about the new community editing paper here.