IGI scientists uncover hundreds of huge viruses with new Cas genes.
In a paper published today in Nature, IGI Investigator Jill Banfield reports a major finding: huge phages containing novel CRISPR systems are ubiquitous across different environments and hosts.
What exactly is a phage?

Bacteriophages, often called “phages,” are an abundant type of virus that infect bacteria and other one-cell organisms. They inject their DNA into a host cell, hijacking the host cell to copy their own DNA and make more phages.
Scientists call the genetic letters that make DNA, “base pairs.” Scientists have long believed that most phages are small in size and have a short genetic code with a small number of DNA base pairs. The average phage genome is thought to be about 50,000 DNA base pairs long. By comparison, the genome of an E. coli bacterium is 1.6 million DNA base pairs, and the human genome is 3 billion DNA base pairs.
Huge phages — everywhere!
“An understudied component of the microbial ecosystem is the predator—which is the phage,” says Banfield. Scientists believe most phages are small, so they usually look for phages with experiments that exclude large particles. To learn more about the roles phages play in different ecosystems, Banfield’s group took a more open-ended approach known as metagenomics. Instead of leaving out larger particles, they sequenced all of the DNA in the samples. This allowed them to find phages of any size. They took samples from different environments, including human saliva, animal feces, freshwater lakes, oceans, hot springs, soil, and material hundreds of meters down in the earth’s crust.
The researchers reconstructed over 350 huge phage genomes, each with over 200,000 DNA base pairs. They put together a 735,000 base pair phage genome—the largest on record. This is more than ten times the reported average size of a phage genome.
The researchers found huge phages in all of the environments they sampled. They infect a variety of different one-cell host organisms. “Large phages have been found before, but they were spot findings,” says study co-first author Rohan Sachdeva. Sachdeva is a bioinformaticist in the Banfield lab. “What we found in this paper is they are essentially ubiquitous. We find them everywhere.” This suggests that huge phages are not an anomaly, but a fundamental part of earth’s many ecosystems.
Secrets of the phage genome

Small phages contain only the most essential genes for using host cells to make more phages. This includes genes that make the protein pieces of the phage body.
“With the larger genome size, you also get a much larger inventory of genes,” explains study co-first author Basem Al-Shayeb, a graduate student in the Banfield lab. They found genes with expected jobs, like redirecting host cells from making copies of their own DNA to making phage DNA or making the host cells burst when they are full of new phage particles. Surprisingly, they also found CRISPR-Cas genes in many of the phages.
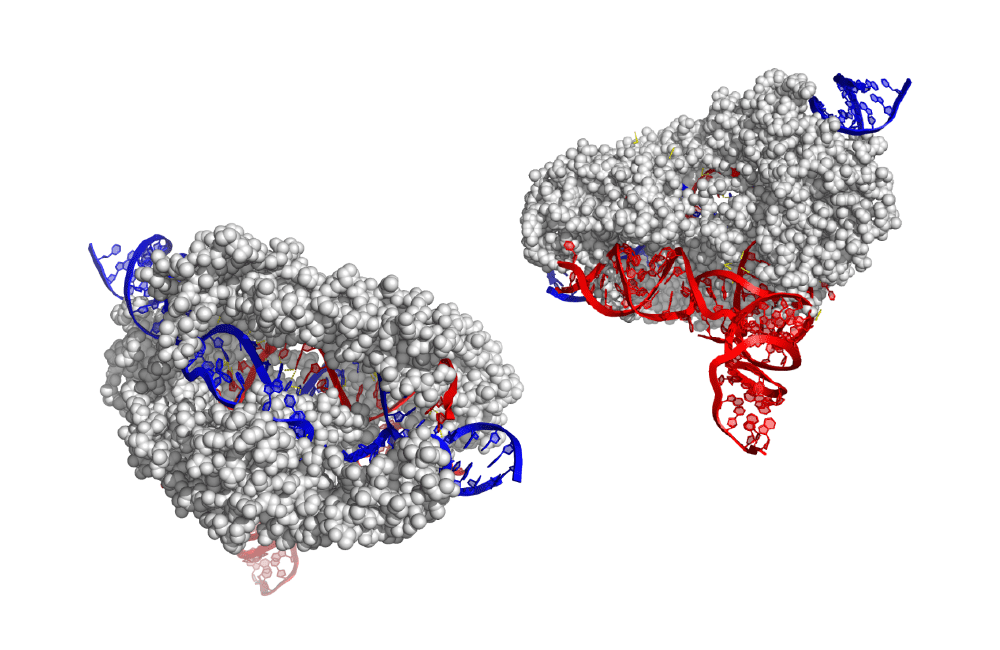
The CRISPR-Cas system evolved as a way for one-cell organisms to defend themselves against phages. In nature, CRISPR-Cas systems recognize phage DNA and cut it up, preventing it from taking over the host cell.
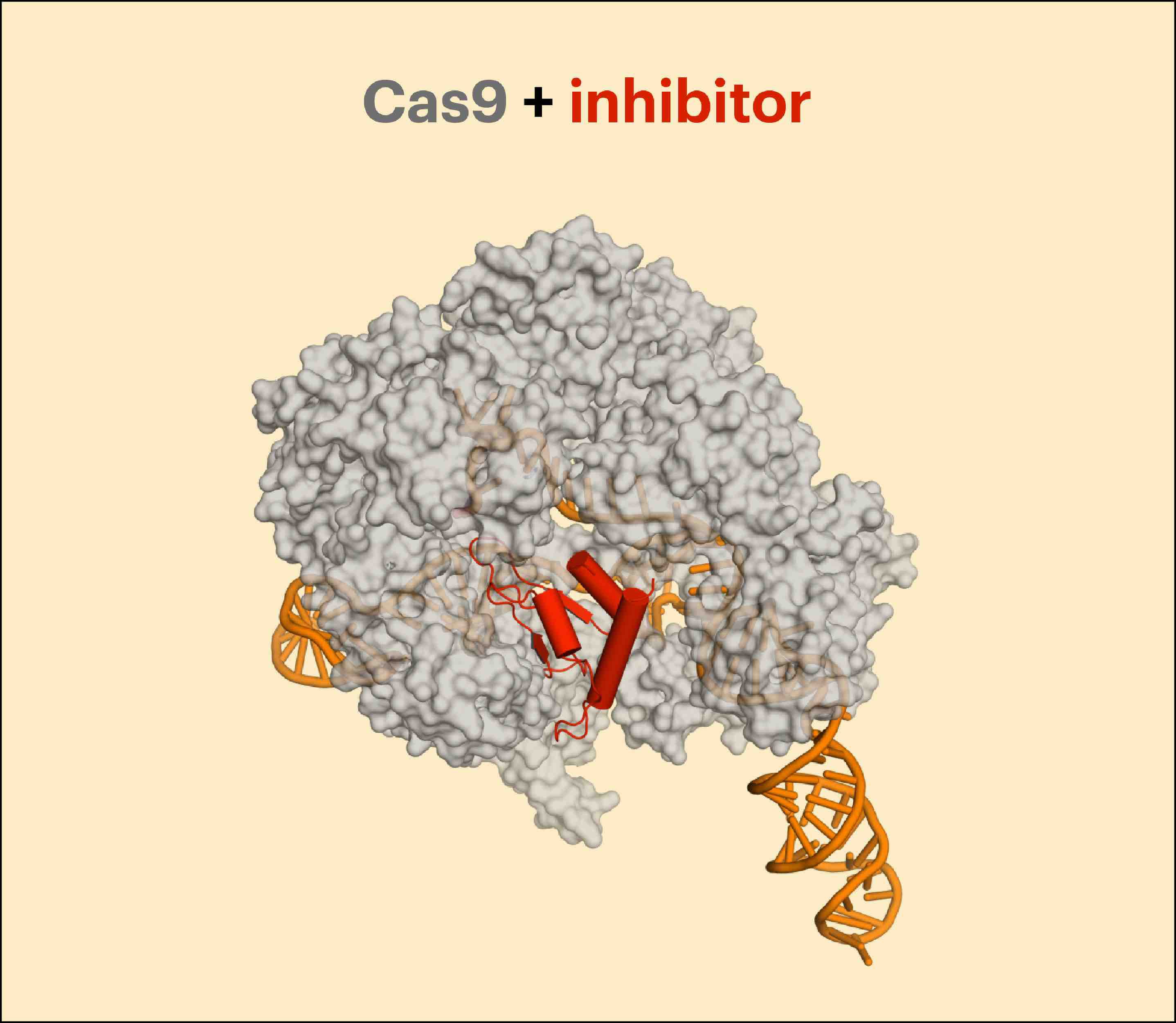
Banfield and her colleagues found the majority of known CRISPR-Cas types in phages, as well as new variants of Cas genes. What is an anti-phage defense system doing inside phage genomes? Banfield suspects that phages use CRISPR to combat other phages that are competing for the same host cell. In other words, a phage uses CRISPR to defend the host cell from other phages—so it can get the host cell resources all to itself. Some phages also contained anti-CRISPRs. Anti-CRISPRs could inhibit host CRISPR-Cas systems that would otherwise cut up phage DNA.
Finding new CRISPR tools

The researchers are now working to grow the huge phages in their laboratories. They want to further study what their genes do. “One of our goals is to access biological diversity in the most comprehensive way possible,” says Banfield. “What are they doing with these genes? The sense we have looking at these large genomes is that phage have acquired a lot of different genes and pathways—some of which we can predict, some of which we can’t—for really taking control of bacterial hosts’ function during infection.”
These phages may also be a new reservoir of CRISPR-Cas tools that could be used for genome engineering. Banfield explains, “Because they are still small compared to bacterial genomes, we expect to find compact and efficient genes which have great appeal for genome editing.”
–
This study was published in the online edition of the journal Nature as “Clades of huge phages from across Earth’s ecosystems.” In addition to Banfield, Al-Shayeb, and Sachdeva, other authors are LinXing Chen, Fred Ward, Patrick Munk, Audra Devoto, Cindy J. Castelle, Matthew R. Olm, Keith Bouma-Gregson, Yuki Amano, Christine He, Raphaël Méheust, Brandon Brooks, Alex Thomas, Adi Lavy, Paula Matheus-Carnevali, Christine Sun, Daniela S. A. Goltsman, Mikayla A. Borton, Tara C. Nelson, Rose Kantor, Alexander L. Jaffe, Ray Keren, Ibrahim F. Farag, Shufei Lei, Kari Finstad, Ronald Amundson, Karthik Anantharaman, Jinglie Zhou, Alexander J. Probst, Mary E. Power, Susannah G. Tringe, WenJun Li, Kelly Wrighton, Sue Harrison, Michael Morowitz, David A. Relman, Jennifer Doudna, Anne Catherine Lehours, Lesley Warren, Jamie H. D. Cate, and Joanne M. Santini.
This research was funded by the Innovative Genomics Institute, University of California, Berkeley, the Joint Genome Institute, Stanford University, Technical University of Denmark, Colorado State University, University of Toronto, Sun YatSen University, University of Cape Town, University of Pittsburgh School of Medicine, Université Clermont Auvergne, Institute of Structural and Molecular Biology at the University College London, the National Food Institute, and the Japan Atomic Energy Agency.
The Innovative Genomics Institute (IGI) is a non-profit, academic partnership between UC Berkeley and UC San Francisco that supports collaborative research projects across the Bay Area. The IGI envisions a world in which genome engineering innovations benefit humanity and are accessible to all. As pioneers in genome editing, functional genomics, and other cutting-edge technologies, IGI scientists continuously push the boundaries of science.
Media Contacts
Megan Hochstrasser: megan.hochstrasser@berkeley.edu
Jill Banfield: jbanfield@berkeley.edu
Basem Al-Shayeb: basem@berkeley.edu