In a paper published today in PLoS Biology, IGI researchers Vivek Mutalik of Lawrence Berkeley National Lab and Adam Arkin of UC Berkeley report advances in understanding phage biology and phage resistance that pave the way towards using these small predators to fight antibiotic-resistant bacteria. In September, Mutalik talked with IGI science writer Hope Henderson about this work.
What does your lab study?
We mainly study synthetic biology and functional genomics of microbes. Primarily, we develop tools to study bacteria, look at them in the microbiome context, and get them to do things. Along the way, I’ve become fascinated by bacteriophages.
Who’s not excited about phages? They’re the coolest things on earth — and we don’t understand them.

They are the “known unknowns” in the microbial world. We want to learn more about phages, how they infect bacteria and how bacteria fight back, and ultimately engineer them to kill specific bacteria for applications in human health and agriculture.
What is a phage?
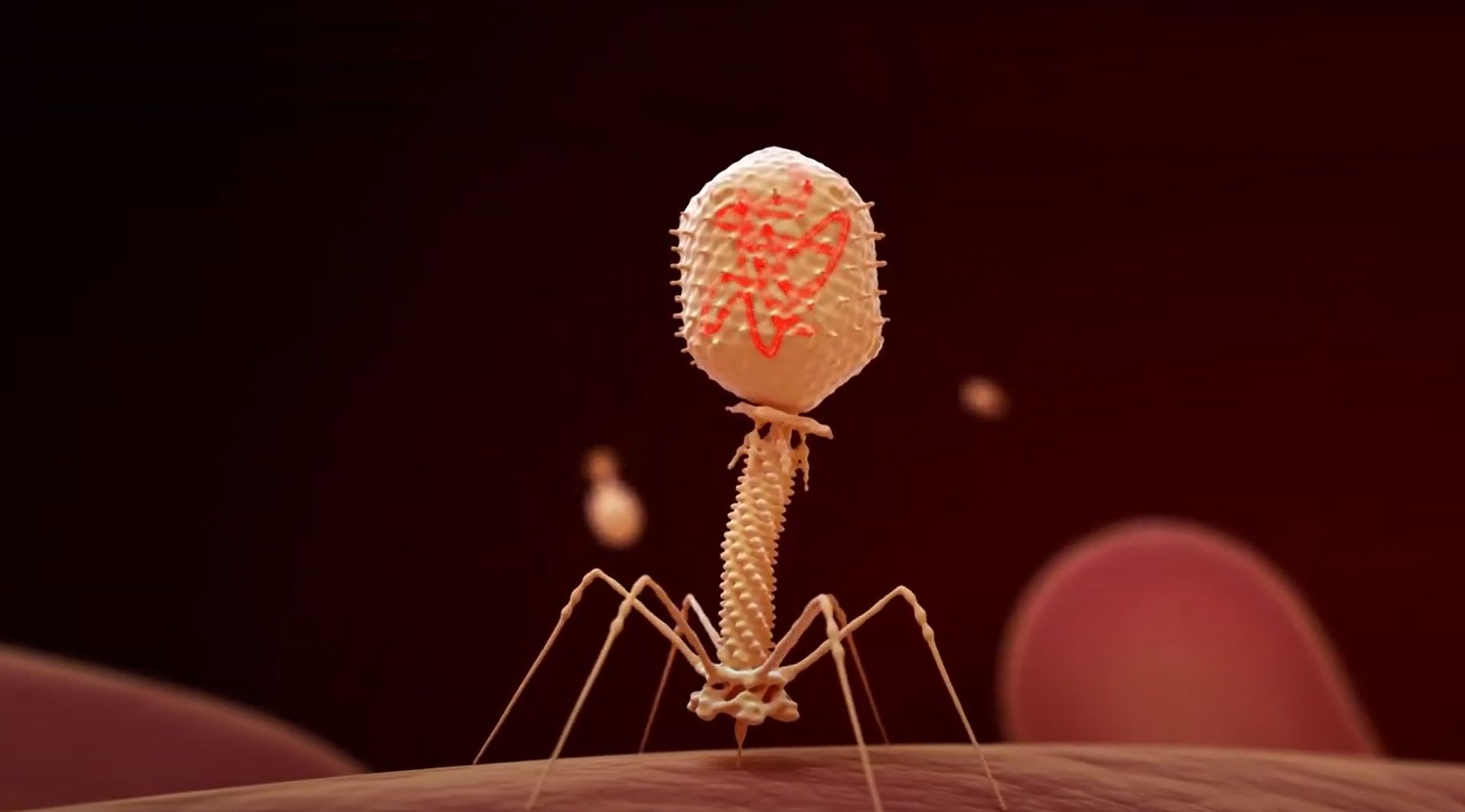
Bacteriophages, or phages for short, are bacterial predators. They are viruses that infect bacteria, and use bacteria to grow and multiply. Phages are found everywhere — they are the most abundant biological entities on earth! Each type of phage has a unique genome, size, shape, and infectivity cycle, of which we have very limited knowledge. But we know that phages are very precise killing machines that each infect specific bacteria. They are not like antibiotics in that when you add antibiotics, a big class of bacteria will be killed. We know that the tail at the bottom of (most) phages determines which bacteria they infect, but we don’t understand exactly how. In addition, even though we are getting better as sequencing phages, we don’t have a good handle on what these phage genomes code for. Essentially, we know who’s there but don’t know what they’re doing.
The ultimate goal is to understand enough phage biology that we can develop approaches for targeted elimination of pathogenic bacteria. Studying phage resistance gives us a window into their inner workings.
What can you do with engineered phages?
The application space is huge: we can use engineered phages with specific “roles” in human health, agriculture, food, meat, aquaculture industry, environmental remediation, and more. If we deeply understand how phages work, we can engineer them to eliminate one particular strain of bacteria in the gut microbiome to help treat an infection, or diabetes, or kill a bacterial pathogen that’s attacking tomato crops.
What does “phage resistance” mean and what does it have to do with engineering phages?
You can think of it as similar to antibiotic resistance. When you add antibiotics to bacteria, they don’t like it! They come up with methods to survive and adapt to the conditions. When you add phages to bacteria, the same thing happens — they go through what we call a selection process. There will be a few bacteria with certain genetic changes that allow them to survive the phages. In other words, the host bacterial population mutates, and now the phage can’t infect them. That’s phage resistance. Studying phage resistance is a way to study the fundamentals of how phages infect specific bacteria. If the changed gene is, say, part of a receptor on the bacteria’s surface, then we know the phage interacts with that part of the receptor to get into the bacteria.

What did you find in this paper?
We used well-studied phages and E. coli, a lab bacteria, to look at phage resistance. Because it’s a known system, we could make sure our platform is effective by recapitulating what has already been found, and we could identify new things.
We took two strains of E. coli bacteria and built libraries of mutations where genes have increased function, loss of function, or are functionally deleted. These libraries are in a test tube or in a flask, and each one has many different mutants in it. Each mutation has a barcode, so to speak, that lets us know what gene is mutated. So then you add a phage. Most of the bacteria are killed, but a few survive and multiply. We collect the survivors and study, why are they surviving? We can use the barcode to tell what mutations they have. The genes we uncover are essential for the phage infection to process. So, I can start to really understand the landscape of phage resistance.
We did that with 14 phages in the paper. Some of these are very well studied, while some are novel phages. Using this system, we could capture almost all of the data on phage resistance that has been generated over 50 years. In other words, the genes we already know are important came up as hits. That lets us know our system is really working and robust. This has huge implications for understanding how bacteria responds to phages and developing ways to engineer phages! The premise of our proposal to the IGI was to show that our platform works for E. coli, and then scale it up to use with a couple of pathogenic bacteria. So now I’m continuing that work with other human and plant pathogens. It’s really exciting to compare these resistance mechanisms across different phages and different pathogens!
Does your work have implications for COVID-19?
Absolutely. Researchers have found that more than 50% of patients hospitalized for COVID-19 also have bacterial co-infections, like bacterial pneumonia. These patients are getting antibiotics, but they don’t always actually kill the bacteria or help the patient — that’s antibiotic resistance. We aren’t giving phages to these patients yet, but they could be part of the arsenal at some point. COVID-19 has shown us that we are just so unprepared for tackling infectious diseases. This is also very important from the national biosecurity point of view especially as we are clearly not prepared to handle any accidental (or intentional) release of antibiotic resistant pathogens into our food processing, water treatment or our environment. Our national strategy seems to be more reactive than proactive. The antibiotic resistance threat has been looming in front of us for so long, but there are neither new antibiotics in the pipeline nor any new solutions being developed. We need a holistic, proactive approach to solve antibiotic resistance, and phages could be a key component of it. I’ve never felt so connected to an application space ever before. I feel very driven.
Whenever I isolate a phage it is with the hope that it is going to save somebody’s life. Maybe not now, but someday. That’s a very satisfying feeling.
Is there anything else you’d like to say?
It was a risky project that is tough to get a funding agency to invest in, and I’m happy that IGI recognized this and took the risk. By investing in this project, IGI has established the expertise, tools and resources needed to do phage characterization and engineering.
I want to say thank you to my team. Specifically, working with postdoc fellow Denish Piya, and graduate scholars Benjamin Adler and Harneet Rishi was super fun. I applied for this IGI grant with Adam Arkin and Adam Deutschbauer as co-PIs — it’s an awesome team to work with these pioneers in functional genomics and synthetic biology! And then having Britt Koskella and Kim Seed on campus, Richard Calender next door, working next to Jennifer Doudna’s lab — it’s just a dream come true.