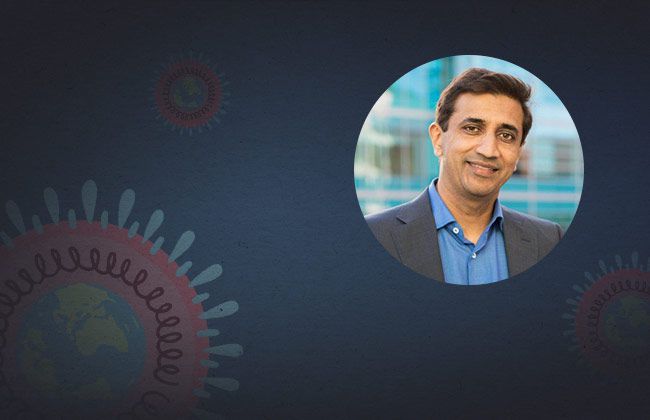
Hani Goodarzi is an Assistant Professor in the Department of Biophysics & Biochemistry, at the University of California, San Francisco. In April, he started working on a research project to uncover the RNA structure of the SARS-CoV-2 genome. In November, he talked with IGI science writer Hope Henderson about this project.
Tell me a little bit about your COVID project.
As you probably know, genetic information is encoded in the molecules A, G, C, and T for DNA. For RNA, it’s A, G, C, and U. SARS-CoV-2, the virus that causes COVID-19, uses RNA instead of DNA for its genetic code. In other words, the RNA molecule is what’s passed along.
DNA is a nice looking molecule, right? It’s a double helix, and the shape is stable. But RNA is just a single strand. For RNA, there is a higher-order level of genetic information in addition to the sequence of letters, which is how this single strand of RNA folds on itself. This is called secondary structure. My team is looking at the secondary structure of the SARS-CoV-2 genome.
How did you get interested in RNA secondary structure?

When I was a postdoc, I started working on a project with Charlie Rice‘s lab, trying to understand whether there is information about when genes are turned on or off or other functional information in the RNA structure of RNA viruses. And actually one of the virus families we looked at were Coronaviruses. So when COVID-19 happened, I already had some inkling of the importance of RNA structure kind in the life cycle of this virus.
How are you looking at secondary structure?
The project we’re working on now is to basically take the viral genome, chop it up, and look at the secondary structures of RNA piece by piece at really high resolution. We’re taking advantage of a protocol that was developed by Jonathan Weissman‘s lab here at UCSF, and we’ve made some modifications to it. And that helps us decode the secondary structure of the RNA genome.
What do you do after chopping up the RNA?
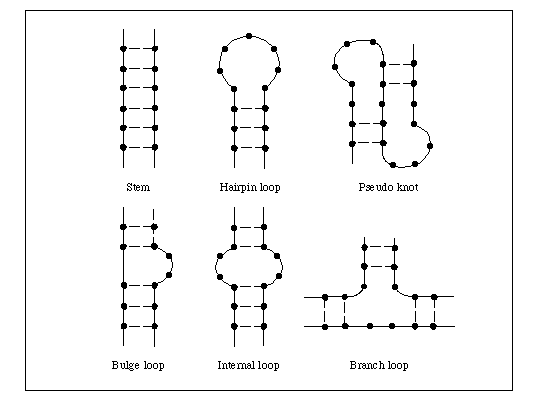
The method is called DMS-MaPseq. The idea is that you add a chemical, DMS, that can modify the RNA. So, how much each RNA nucleotide — what we call an individual A, C, G, or U — is modified depends on how exposed that nucleotide is. For example, if the RNA is folded into itself or forming something like a double helix, the chemical can’t access it anymore and you won’t see a modification. If it’s in, say, a loop or a bulge where it’s open, you will get modification.
On top of this chemical reaction, you add a high-throughput sequencing methodology that reads out these modifications as mutations in the sequence. The more mutations you see after you add this chemical, the more modification there was, and the more exposed that nucleotide was. You can combine the data on which nucleotides were mutated with the RNA sequence and make predictions about the secondary structure of the RNA.
Will understanding the SARS CoV-2 viral structure and the meaning of that structure have implications treating COVID-19?
Yes. When we talk about viral RNA being functional, it means that they play a role in the life cycle of the virus. And there are a couple of ways that you can target them therapeutically. One is to target the RNA structure itself. It’s unusual, but there are a few pharmaceutical companies that are really focused on targeting RNA itself and stopping the cell from using the RNA to make proteins.
Another way knowing structure can be used to create therapeutic targets is based on the fact that RNA structures can be a hub of interaction, where other crucial factors act. You have RNA-binding proteins that interact with it. You have helicase enzymes that open the RNA up.
So, if you figure out the network of proteins that are interacting with the viral genome, you can target and disrupt those interactions as well. For example, if a helicase is needed to open the RNA structure for the replication of the virus, then inhibiting that helicase should bring down the viral replication.
Where are you in the project?
The first step of the project was to basically chop up the genome and then clone the individual pieces into bacteria that we use as factories to make more copies of the genome piece. So that’s where we are. We’re troubleshooting this part. Next, it’s crucial to understand how the RNA actually acts in a human cell. So we want to put the RNA fragments into human cells, and then use DMS-MaPseq technique I mentioned.
Are these shorter, “chopped up” pieces of RNA an accurate representation of how the RNA forms a secondary structure when it’s in one piece?
There are some papers that have already solved the RNA secondary structure of SARS-CoV-2, but with lower resolution than what we intend to do. But that means we have some information about the longer range interactions that we can work off of. We will probably miss some things by using fragments, but this will allow us to do this at a higher resolution. So it’s a pros vs. cons scenario.
Do you have any new quarantine hobbies?
We bought an indoor hydroponic farm and started growing herbs.
How are your lettuces coming out?
They’re great! So are the cherry tomatoes and herbs. Highly recommended!
This work is funded by the Laboratory for Genomics Research (LGR), a collaboration between UC Berkeley/UCSF (IGI) and GlaxoSmithKline.



