Meet an IGI Scientist: Benny Ordoñez
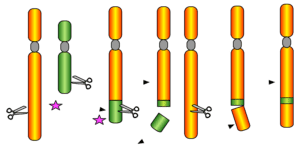
This series introduces the public and fellow researchers to our talented scientists. We interview different IGI members to find out who they are and what makes them passionate about science. — Benny Ordoñez is a graduate student in the Comai lab at UC Davis. Her work harnesses the genome editing power of CRISPR-Cas9 technology for […]