
Bruce Wang, M.D., is an Assistant Professor in the School of Medicine at UCSF. He is a PI on an IGI-funded research project looking at the effect of SARS-CoV-2 infection in different tissues. In April, he talked with IGI science writer Hope Henderson about this project and how it’s progressing.
Tell me about your COVID-19 project.
 Our project is in trying to understand how SARS-CoV-2 affects different organs in severe COVID-19 disease. This is a collaborative project amongst six different labs at UCSF and the Chan Zuckerberg Biohub. Before this, we were all involved in projects to characterize the cellular makeup of each tissue — a cell atlas type of project, if you will. Each one of our labs has expertise in a particular mammalian organ, so the collaboration made a lot of sense.
Our project is in trying to understand how SARS-CoV-2 affects different organs in severe COVID-19 disease. This is a collaborative project amongst six different labs at UCSF and the Chan Zuckerberg Biohub. Before this, we were all involved in projects to characterize the cellular makeup of each tissue — a cell atlas type of project, if you will. Each one of our labs has expertise in a particular mammalian organ, so the collaboration made a lot of sense.
What do you hope to learn by looking at tissues from patients with COVID-19?
We are interested in two main questions: (1) Can we figure out what cells SARS-CoV-2 targets within each one of these organs? (2) Is the virus directly affecting them, or are effects in different organs part of the body’s systemic response to infection in the lungs?
And, what are the changes that are happening in each organ? Are the kinds of cells in each tissue changing? We were especially interested in whether the number or proportions of kinds of immune cells are changing in these organs.
What approaches are you using to answer these questions?
We are trying to then map all of the cells that we can identify in the tissue to get an idea of their spatial relationships with each other — this gives us a lot of information about what is happening in an organ, for instance if immune cells are invading a certain part of an organ.
And, we are capturing individual cells from each one of these organs and looking for the presence of the virus, as well as measuring their transcriptomes using single-cell RNA sequencing. The transcriptome tells us what kinds of things the cell is trying to do, by telling us what proteins it is actively making RNA for.
How do you get tissue samples for your work?
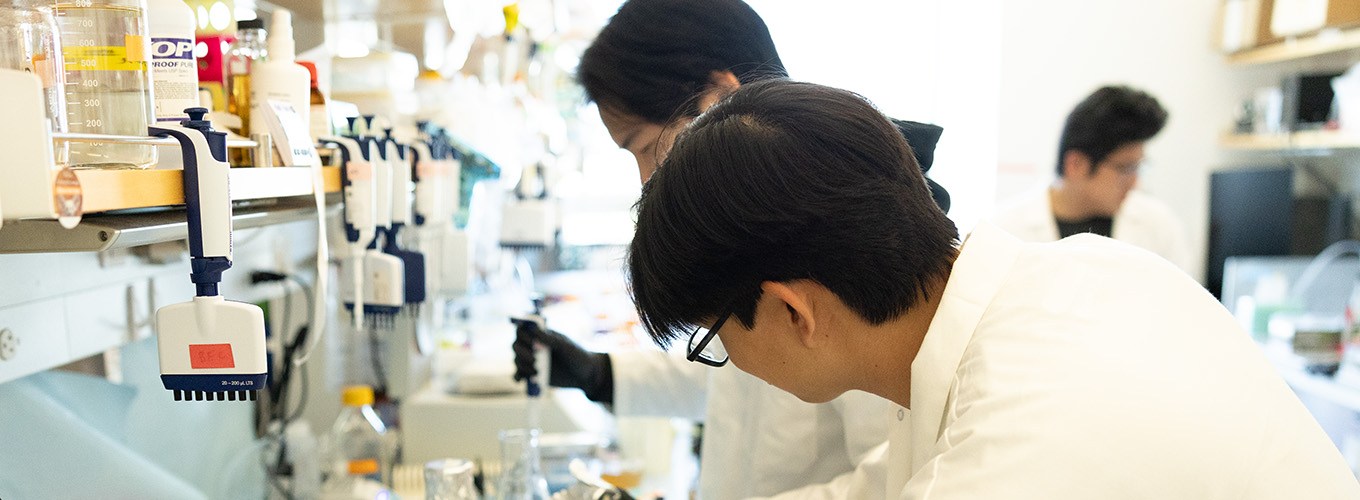
This project was really only possible because we had pathologists who were very willing to help and collaborate. We get our tissues from two different sources. One was locally at UCSF where the pathologist set up a COVID-19 autopsy service. So, there was a separate room that was set aside where the pathologist would work in extensive personal protective equipment. The pathology department got institutional review board approval to get de-identified human tissues, meaning we do not receive information that could be used to tell who the sample belonged to, so anonymity of the deceased person is protected.
We also got samples from a collaboration with Saarland University in Germany, who had a similar type of autopsy service set up there that they were very generous with. We are really grateful to the patients and families who allowed the autopsy and tissue collection. Nothing can make up for losing a loved one, but hopefully there is a little solace in knowing that they are contributing to research on understanding and fighting this disease.
Did you find evidence of the virus in different tissues?
In the vast majority of the tissue samples, we actually can’t detect any evidence of the virus. We have limited information on patients, but we do know they were in the hospital for an average of close to four weeks before they passed away. We think what has happened is that the acute COVID infection was early on in their disease course, and then their body probably cleared most of the virus. And by the time they passed away, it was probably the intense immune reaction to the illness, along with lung dysfunction, that led to multiorgan failure.
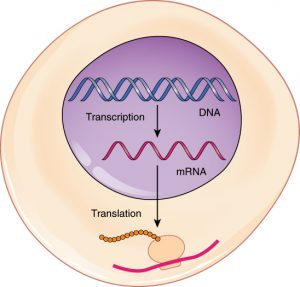
We weren’t able to get sufficient numbers of high-quality live cells from the tissue sample, likely because these are postmortem tissues, but we were able to capture intact nuclei — the compartment of the cell that holds DNA, and where you can find mRNA right after it’s made. We’ve been sequencing the mRNA from the nuclei in different tissues; so far, we’ve processed about 50 organ samples and looked at over 250,000 single nuclei, each representing a single cell. We only need a very small amount of tissue to do this.
What can you learn from looking at nuclei?
Within these samples, we compare the RNA from the small number cells we found that are positive for SARS-CoV-2 with other cells in the same tissue. We can also compare the RNA from patients with COVID to those from patients that had no infectious disease, or who died from influenza. We thought this was an especially interesting control, since they had a different viral infection leading to lung disease.
It’s clear that severe COVID infection leads to widespread changes in what genes are active in a cell. We’re using an approach known as principal component analysis to get a sense of the changes. And COVID infection status consistently comes out as the first principal component, in other words, it explains the most variance in our dataset.
What are you looking forward to as people get vaccinated and restrictions start to lift?
This past year of Zoom meetings and discussing data over Zoom is nowhere near as productive or as engaging as when it can be done in person. What I’m looking forward to is when we can get to that point where infection in the community is so low and everybody’s vaccinated and we can start having more face-to-face discussions.



