Genomics
Institute
A high-throughput functional genomics platform to combat fungal growth and mycotoxin contamination of food supplies
We are developing new genetic tools to combat fungal food contamination.
SHARE:
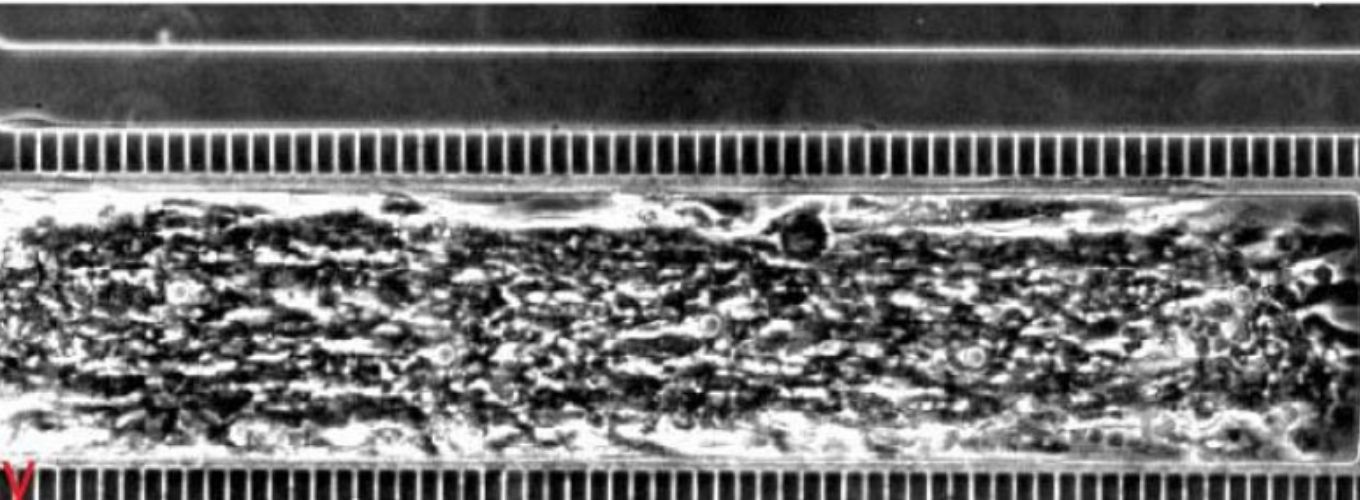
Modern genome-editing tools allow biologists to easily modify an organism’s genome. However, scientists currently do not know the function of the vast majority of genes. In order to fully exploit our genome editing capabilities, we will need to rapidly increase our knowledge of gene function. Recent advances in high-throughput screening methodologies have allowed us to decipher gene function and define novel pathways. While these tools work well for single-celled organisms, such as yeast and bacteria, many of the fungal species responsible for devastating crop infestations that threaten global food supplies grow as filaments, with multiple nuclei in a single cell. These differences in growth habit make simple transference of high-throughput technologies developed for single cells to filamentous fungi a challenge.
Our overall goal is to develop high-throughput loss-of-function and gain-of-function libraries in two filamentous fungal species: the model filamentous fungus Neurospora crassa and the toxigenic mold Aspergillus flavus, which infects key staple crops and produces toxins that are both acutely toxic and carcinogenic. These libraries of mutants will be subjected to high-throughput assays that test thousands of conditions for fitness data collection and analyses. This approach will lead to the development of a new suite of tools to interrogate gene function in filamentous fungi and new strategies for mitigating A. flavus contamination of food supplies, thereby improving human health and ensuring food security.
Share this project: