Metagenomics 101 With Spencer Diamond
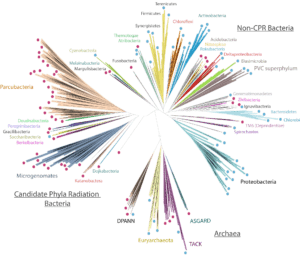
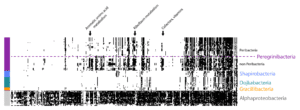
Jill Banfield and her team have used genome-resolved metagenomics to find new CRISPR systems, huge viruses, whole new branches on the tree of life, and more — but what exactly is metagenomics? We wanted to take a step back and introduce readers to the Banfield lab’s pioneering techniques for finding and understanding new microbes, with […]