Genomics
Institute
Engineering CRISPR-guided DNA polymerases for in vivo, unbiased mutagenesis of user-defined loci
We are developing versatile technology for rapid “directed evolution” of desirable traits in laboratory organisms.
SHARE:
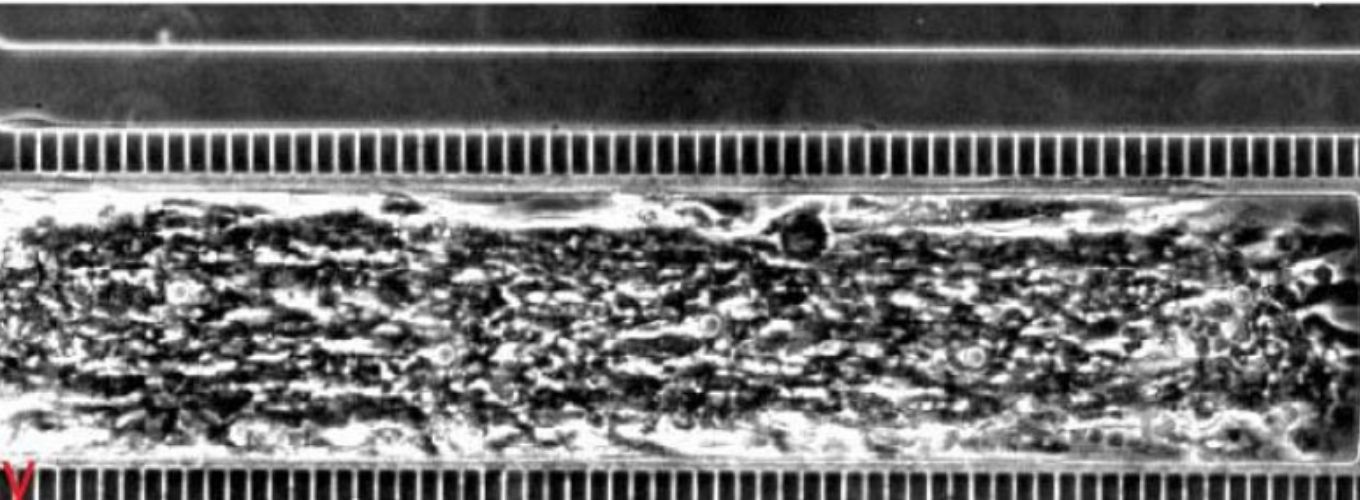
Living organisms have evolved incredibly complex and useful traits by sampling immense sequence diversity under selective pressures over many generations. In the laboratory, where both diversity generation and selective pressures can be manipulated, artificial application of natural selection - referred to as “directed evolution” - has emerged as a powerful method for isolating genetic material with desirable properties from a library of sequence variants. However, the sequence space that can be explored is limited by the efficiency of synthesizing and transforming a genetic library. The requirement for efficient transformation rates has especially confined directed evolution to only a few model organisms.
The ability to program a cell to elevate mutagenesis at localized, user-defined loci would remove the need to transform a synthesized library of nucleic acids; however, current in vivo targeted mutagenesis platforms are unfortunately either confined to targeting a particular, hardwired locus in a specific organism (e.g. a phage genome, or the IgG locus) or have a narrow and biased editing window at user-defined loci. We are developing the first unbiased targeted mutagenesis system capable of evolving phenotypes encoded by large stretches of user-defined loci. We anticipate that this crosscutting technology will be further generalizable to other microbes and higher-organisms (such as plants).
Share this project: