In nature, CRISPR is an immune defense that bacteria and other microbes use to protect themselves against viruses by recognizing and cutting the genomes of invading viruses. In 2020, researchers at the Innovative Genomics Institute found a CRISPR-Cas system in what would seem to be an unlikely place: inside a virus.
In a new paper in the journal Cell, a collaboration between multiple IGI investigators at UC Berkeley including Jennifer Doudna, Jill Banfield, David Savage, and Brian Staskawicz, as well as Steve Jacobsen’s lab at UCLA, reports new findings that the diversity of CRISPR systems in viruses that infect bacteria (bacteriophages or “phages” for short) is far richer than expected and could be a valuable source of new, efficient genome editors for use in humans, plants, and other organisms.
Good CRISPR Hunting
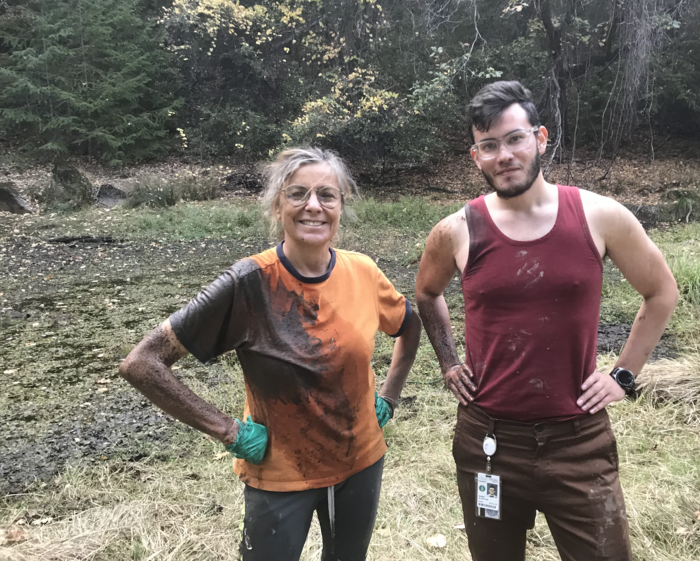
Finding CRISPR systems in phages was not a total fluke. First author Basem Al-Shayeb, who was a graduate student in the Banfield and Doudna labs when this research began, had a reasonable suspicion that he would find CRISPR systems in phages and that they could prove useful for genome editing.

“Phages tend to have more constraints on genome size than a lot of bacteria, so the proteins that are encoded on their genomes tend to be much smaller. I reasoned that phages could be a very good place to find compact CRISPR-Cas systems,” says Al-Shayeb.
For gene editing, size matters: the smaller the gene editing molecule, the easier it is to deliver it into cells so that it can make the desired changes to the DNA. In the 2020 study, the team found CRISPR systems in a group of unusually large phages called “megaphages.” While the phages were huge – that is, for viruses – the CRISPR-associated proteins responsible for actually cutting DNA, known as “Cas proteins” for short, were tiny relative to the widely used Cas9 protein from bacteria. And the new protein wasn’t just small, it worked: the team was able to adapt this new protein called CasΦ (Cas-phi) for gene editing in mammalian cells and in plant cells. That find was just the tip of the iceberg.
For the new paper, Al-Shayeb and colleagues conducted a broader survey across bacteriophages using metagenomic analysis of soil, aquatic, human, and animal microbial samples and found an unexpected diversity including all six known types of CRISPR systems spread across over 6000 types of phages, not just megaphages. Like CasΦ, these newly identified Cas proteins were all super compact compared to those found in bacteria.
Antiviral Viruses
One key question is why are viruses walking around with CRISPR systems… that kill viruses? As a rough comparison, imagine your surprise if you found an ant armed with a tiny can of Raid.
“That was my first question,” says Abby Stahl, a postdoc in the Doudna lab and co-author on the paper. “I thought that CRISPR systems were predominately used by bacteria for anti-phage defense, and suddenly I’m learning that they’re also found in bacteriophages. What are they doing there?”
The leading theory is that these CRISPR systems give phages an advantage when there’s competition from other phages: the invading phage gets a host to itself, while its CRISPR system helps defend against other would-be attackers. How different lineages of phage were able to acquire CRISPR systems at all remains unanswered.
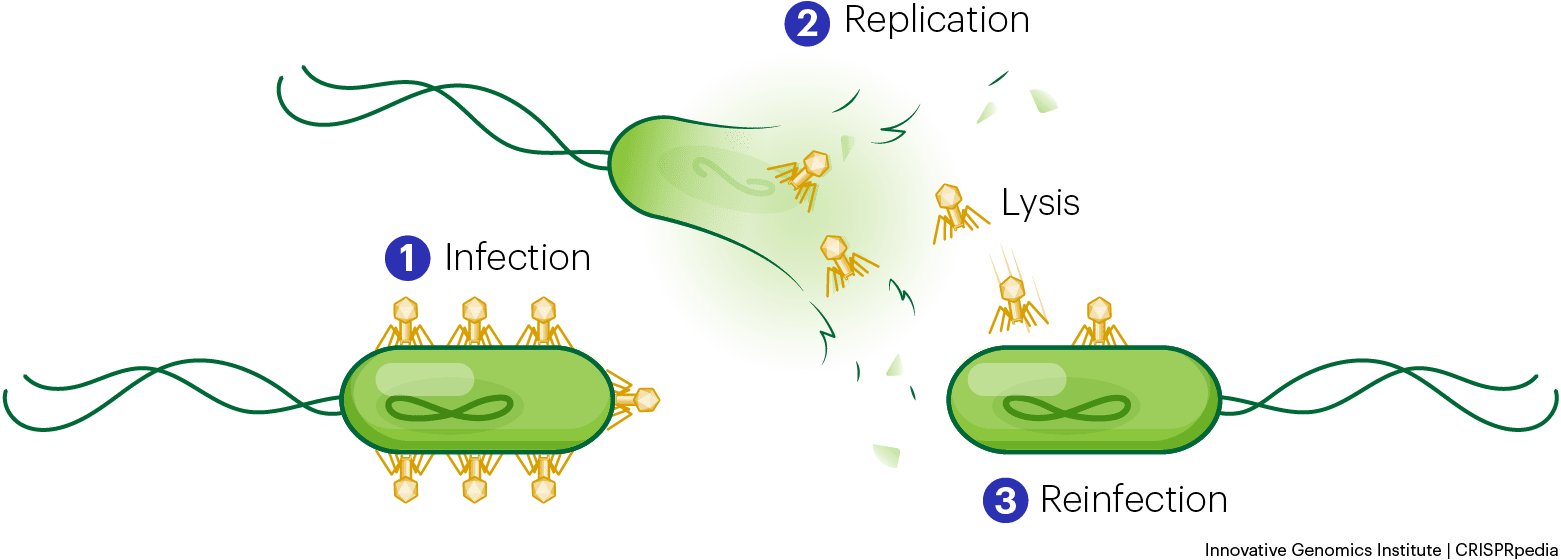
What is clear is that these phage-encoded CRISPR systems are not just small, they’re unique. Among the diversity of newly described CRISPR systems and Cas proteins, the team chose to focus on a particularly small protein with structure unlike any other previously described, which they call Casλ (Cas-lambda).
“Speaking of the protein itself, its structure is distinct from any known Cas protein, including the other viral hypercompact nuclease CasΦ,” says Petr Skopintsev, a postdoc in the Doudna Lab who delved into the structure of Casλ using cryo-electron microscopy. “When you’re looking for new CRISPR systems, you’re looking for something that doesn’t look like what has already been identified. This protein was distinct. It’s a new type of hyper-compact protein that has good activities for genome editing.”

To verify this experimentally, the team tested Casλ in the lab to edit the genomes of mammalian and plant cells and compared its efficiency against Cas12a, the most closely comparable widely used genome-editing Cas protein. Without any additional engineering, Casλ performed well across the different cell types.
“We benchmarked it against Cas12a, which is a gold standard in the field like Cas9. The fact that it can perform like or better and be so small is a huge advantage for treating genetic diseases. But we also edited multiple different plants including wheat, which is traditionally very difficult to edit,” says Al-Shayeb.
Even with this broader survey, the hunt for better gene editors and other useful genomic tools is far from over.
“There’s always more to explore,” says Al-Shayeb. “That’s the fun of it.”
Read more:
Basem Al-Shayeb, Petr Skopintsev, Katarzyna M. Soczek, Elizabeth C. Stahl, Zheng Li, Evan Groover, Dylan Smock, Amy R. Eggers, Patrick Pausch, Brady F. Cress, Carolyn J. Huang, Brian Staskawicz, David F. Savage, Steven E. Jacobsen, Jillian F. Banfield, Jennifer A. Doudna. 2022. Diverse virus-encoded CRISPR-Cas systems include streamlined genome editors. Cell 185 (24): 4574-4586. https://doi.org/10.1016/j.cell.2022.10.020.
- Megaphages harbor mini-Cas proteins ideal for gene editing (Berkeley News)
- CRISPR is so popular even viruses may use it (Science)
- CRISPR tools found in thousands of viruses could boost gene editing (Nature)
Media contact: Andy Murdock, andymurdock@berkeley.edu