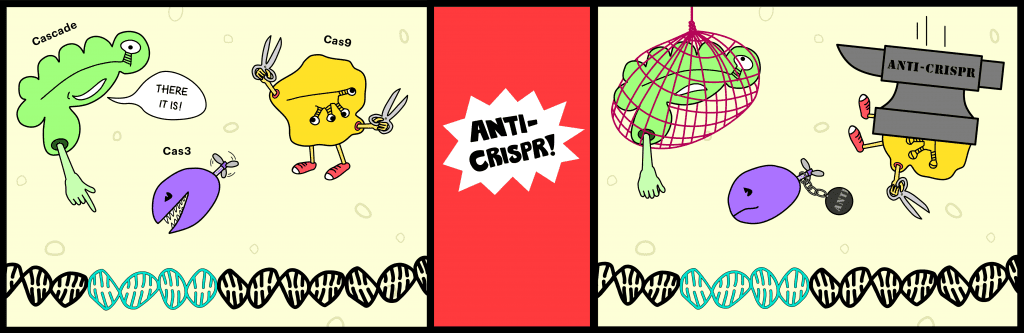
While human wars can last hundreds of brutal years, they’re nothing compared to the ongoing battle between bacteria and the viruses that infect them, bacteriophages. This bitter feud has stretched on for hundreds of millions of years, fueling a so-called “evolutionary arms race.” Bacteria have built various molecular reinforcements to push back the viral onslaught, but all these defenses can be overcome. Eventually, the prokaryotes evolved an adaptive immune system: CRISPR. Like its human counterpart, this potent pathway lets bacteria remember past threats and quickly quash returning viral invaders. You might think the story would end there, but phage struck back with a fascinating arsenal of “anti-CRISPR” proteins.
Seek and ye shall find
The first CRISPR-Cas system inhibitors were revealed in January 2013, around the same time that Cas9-mediated genome editing was successfully demonstrated in human cells. Alan Davidson and his lab were studying bacteriophage biology at the University of Toronto and wondered why phage had been able to overcome so many bacterial defenses but couldn’t seem to circumvent CRISPR. Along with phage expert and frequent collaborator Karen Maxwell, they predicted that phage-encoded “anti-CRISPR” activity might exist and set out looking for it.
The hunt paid off, yielding five distinct families of phage genes that inhibit the I-F CRISPR system in Pseudomonas aeruginosa. After this landmark discovery, the same group found more and more genes encoding anti-CRISPR proteins, 14 families in all. So far, all of the identified proteins are tiny, only about 50–150 residues long (~10–20 kDa). Many of the proteins block the I-F system, others target I-E, and one, co-discovered with Peter Fineran, actually inhibits both! Unlike the streamlined Type II system, with its single enforcer, Cas9, Type I systems employ a multi-protein interference complex called Cascade to bind DNA targets and recruit the nuclease-helicase Cas3 to destroy them. Effector complexes from different subtypes possess distinct subunits with extremely low sequence homology, so cross-subtype inhibition by an anti-CRISPR was unexpected and intriguing.
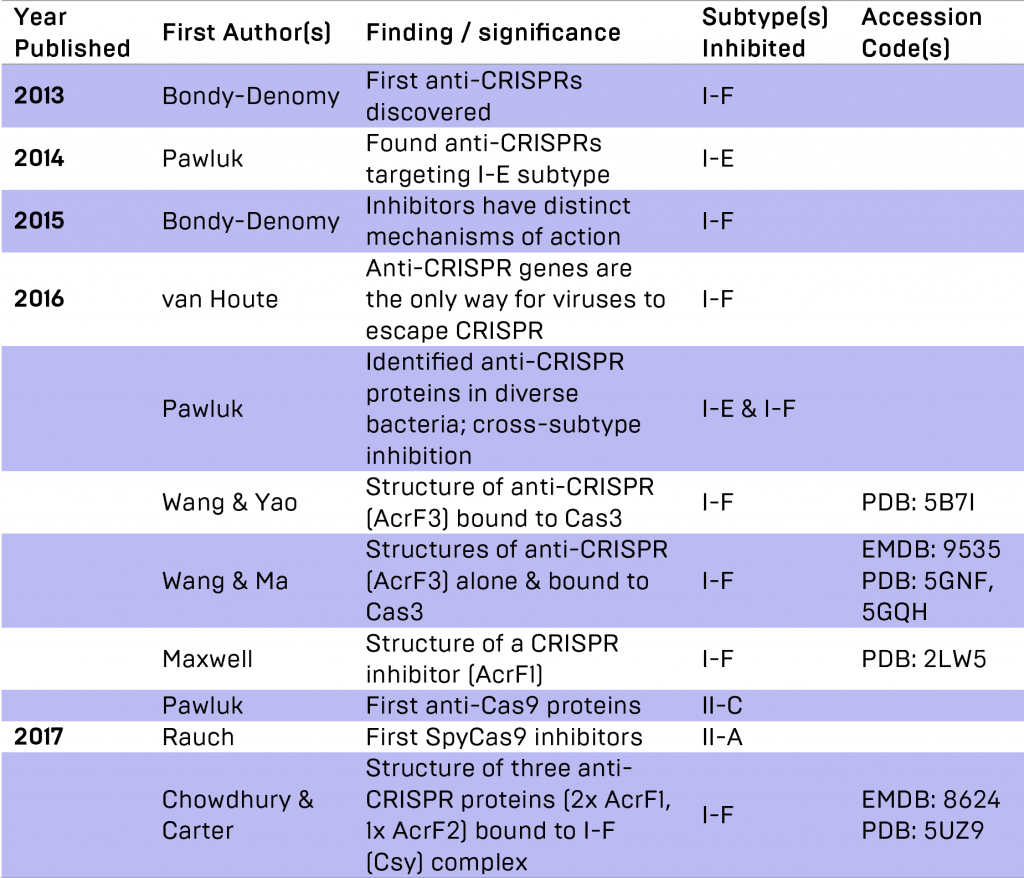
Summary of anti-CRISPR publications to date.
Puny proteins pack a punch
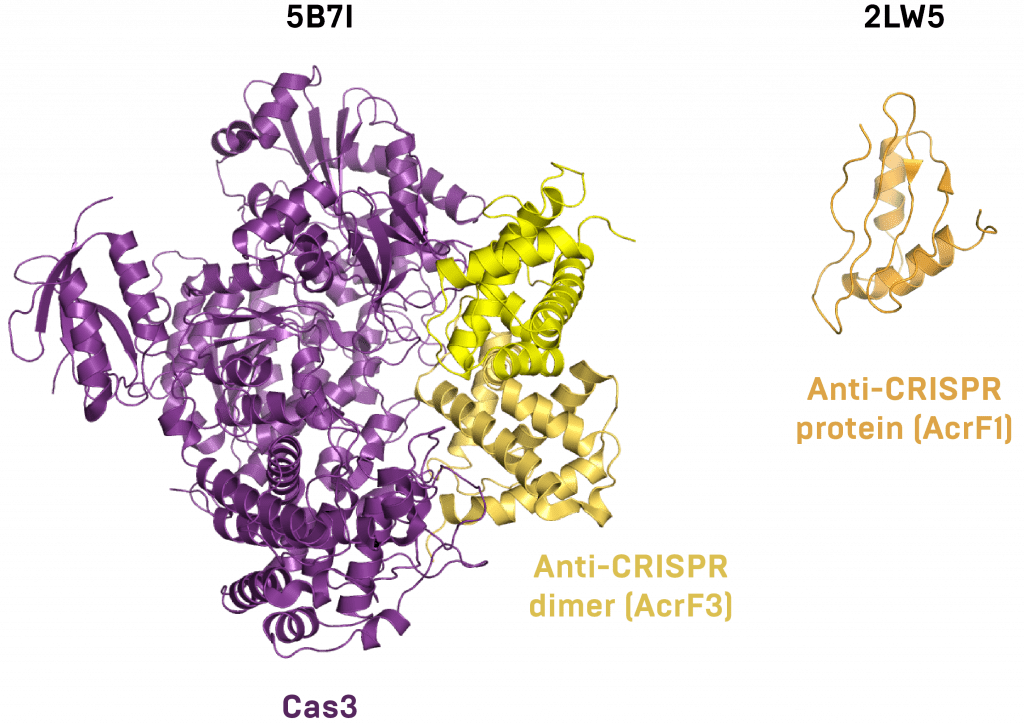
This finding came off the heels of another mechanistic surprise. A year before, the Canadian team published an incredible paper showing how each of the identified I-F inhibitor proteins gums up the CRISPR works in a unique way. They collaborated with Blake Wiedenheft, an expert on the I-F interference complex and a central player in CRISPR biology research. Together, they discovered that two of the anti-CRISPRs block the I-F complex from binding DNA, but they bind to different sets of subunits. Another sticks to Cas3 and prevents its recruitment to the I-F complex.
This interaction was further detailed in summer 2016, when a new group entered the fray and crystallized the inhibitor-Cas3 co-complex. This anti-CRISPR seems to act as a dimer, preventing Cas3 from hydrolyzing ATP, binding DNA, or contacting the interference complex. Another group of newcomers published a similar structure and one of the anti-CRISPR alone just a few months later. Next, Maxwell and Davidson’s groups published an NMR structure of a different anti-CRISPR protein and mapped its functional interface.
Atomic structures of anti-CRISPR proteins bound to the Cas protein they inhibit (left) or alone (right).
These structure-function studies indicate that anti-CRISPR proteins are versatile, powerful weapons, especially when a phage carries more than one or co-infects with another well-armed prokaryotic virus. Even if Cas proteins evolve to mask the binding surface of one inhibitor, another anti-CRISPR widget will still do the trick.
All together now
Following up on their fruitful collaboration, Davidson, Maxwell, and Wiedenheft teamed up with cryo-electron microscopy wiz Gabe Lander and embarked on structural studies. In a tour-de-force paper released online at the end of March 2017, the crew determined an atomic model of the I-F interference complex adorned with three anti-CRISPR proteins. Not only is this the first high-resolution structure of the I-F effector complex, it is also the first structure to show how multiple inhibitors can simultaneously bind and cripple a CRISPR complex, providing fascinating mechanistic insights.
One of the anti-CRISPRs prevents the complex from binding target DNA by sticking to two of the repeating backbone proteins. Because there are six of these subunits, the structure accommodates two separate copies of this inhibitor, and their previous work suggests that up to three may be able to bind simultaneously. To me, the most remarkable finding is that the other anti-CRISPR protein appears to work by elegantly mimicking double-stranded DNA (see below). Read the full paper here.
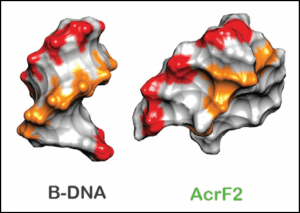
Negative charges (red and orange) in B-form DNA and in the anti-CRISPR protein AcrF2, where they are arranged in a pseudo-helical pattern.
Gene editing with safety scissors
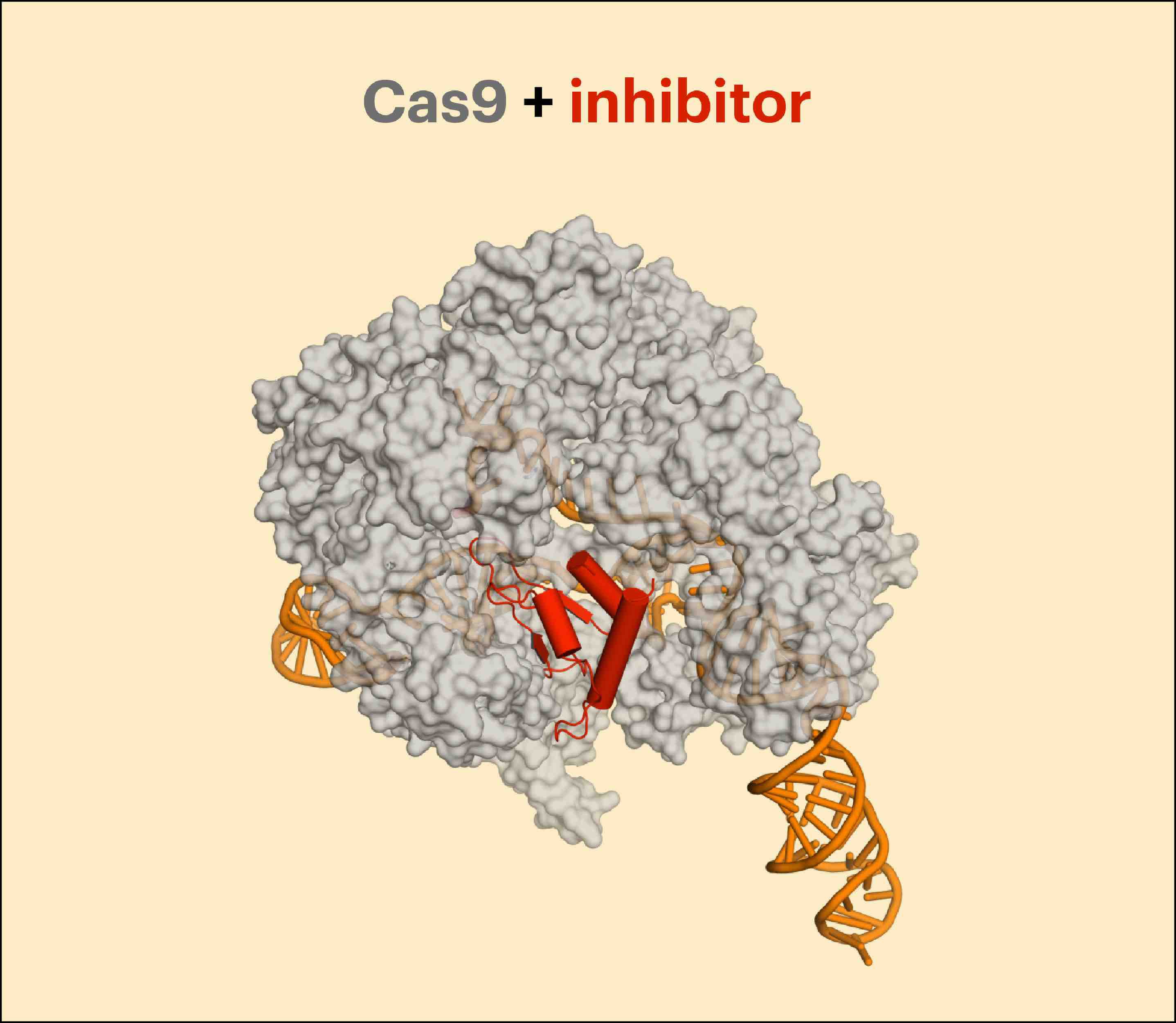
In addition to the exciting progress characterizing Type I CRISPR inhibitors, the first anti-CRISPR proteins that hinder a Type II system were unveiled at the end of 2016. Along with Erik Sontheimer, RNA biologist and CRISPR innovator, the Toronto team found three new families of anti-CRISPRs that specifically inhibit the Type II-C Cas9 protein from Neisseria meningitidis. The inhibitors bind directly to Cas9 and can be used to block both genome editing and transcriptional repression via CRISPRi in cultured human cells.
Less than weeks into the new year, UCSF Faculty Fellow and Davidson lab alum Joe Bondy-Denomy described four more anti-CRISPRs targeting Type II systems. Excitingly, these newly discovered proteins inhibit the type of Cas9 most widely used for gene editing, the Type II-A effector from Streptococcus pyogenes. Accordingly, these anti-CRISPRs can be used to diminish both genome editing and gene repression in human cells.
There’s a neat video abstract to complement the 2016 paper, check it out below:
What’s next?
Researchers have identified 21 families of anti-CRISPR proteins to date. I don’t have the space to get into the interesting approaches used to find these novel inhibitors, but I expect that more and more anti-CRISPRs will be discovered through these and future methods. I hope to see inhibitors of Type III systems, which employ the Csm and Cmr complexes, as well as Class II single effector proteins other than Cas9: Cas12a (Cpf1), Cas12b (C2c1), Cas13a (C2c2), Cas13b, CasX, CasY, and whatever else is coming down the pike.
Beyond the interesting biology, Cas9 inhibitors are welcome additions to the genome manipulation toolbox. While scientists have concocted many inducible Cas9 schemes, there had been no off-switches. Anti-CRISPRs add another layer of control to genetic engineering. They could potentially be used to avoid off-target edits, inactivate cutting in non-target tissues, or block activity at inappropriate cell cycle or developmental stages, particularly to prevent editing events that cause undesired mosaicism in animal embryos. CRISPR-Cas9 off-switches may be especially important in ongoing discussions and research toward responsible gene drive design and biosecurity.
I’m sure there are many applications I can’t even imagine. How well the in vitro and cell culture work translates to usage in a living organism remains to be seen, but it will be great to see what people come up with. Anti-CRISPRs have already made their way into science fiction plotlines, who knows where they’ll end up next?